October 4, 2017 | Core Module
Arkgene: Perfect for Microbial Genomics

The science of microbiology was dramatically changed when the complete bacterial genome of Haemophilus influenzae was first published in 1995. Since then, more than 30,000 bacterial genomes have been sequenced and publicly available in NCBI, with majority of them from pathogens, a testament to how much advances had been mad in DNA sequencing technology.
The pathogenic bacteria are receiving utmost attention in microbial genomics research, with the research objectives to study microbial diversity and population, virulence, symbiosis, drug susceptibility and disease resistance. Extraction and interpretation of these biological information from genomic data is predominantly important so that infectious diseases can be managed, controlled and eradicated more effectively to safeguard human health.
The Arkgene team, in working towards further development in the study of microbial genomics, started Arkgene for Microbes, intentionally designed to provide essential computing power, storage and analysis tools in order to assist researchers in microbial genomic research. Researchers have routinely faced problems in server/hardware setup, running the command line execution, Linux and UNIX like operating systems, and complex bioinformatics algorithms in their work. With Arkgene’s graphical user interface (GUI)-based platform, researchers will be able to have much improved efficiency in their data management.
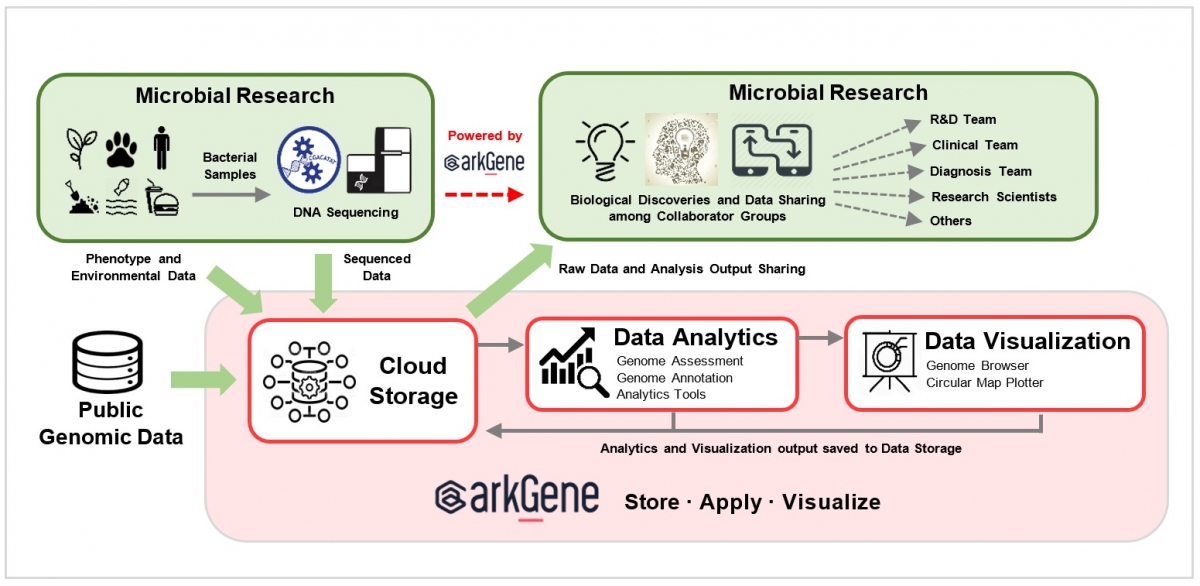
Research institutions, medical centers and diagnosis labs can upload and store their sequenced microbial genomic data in the Arkgene cloud platform, accelerating the secure data sharing among global researchers and networks with permission-based controls. Our Microbial Genome Assembly Completeness Assessment Pipeline can assess the completeness of the bacterial genomes and generates output results in an interactive graphical form. The gene structures (e.g. tRNA, rRNA and protein coding gene) and functional annotation can be generated in one click and a few minutes by using our Microbial Genome Automated Annotation Pipeline.
Our experienced bioinformatician had optimized the two above-mentioned automated analysis pipelines to output the optimal results which can be seamlessly viewed using a set of visualization tools in Arkgene. For instance, the annotated bacterial genome with structural information can be visualized as circular map plot by using our ClicO FS (doi.org/10.1093/bioinformatics/btv433) integrated into Arkgene. Besides, it can also be co-visualized with other genetic information such as DNA markers and gene expression data by using our CGBase genome browser.


In addition, Arkgene offers a set of comprehensive tools for downstream analysis of microbial genome, such as BLAST search against customized database (e.g. virulence factor and antibiotic resistance databases) and ePCR to check the specificity of designed primers.
With the creation of Arkgene for Microbes, not only molecular microbiologist will get access to better tools for their work, but it will also be greatly beneficial to any scientists who works with microbial data.
